Software
We advocate for open methods and software in computational biology, and we’re serious about writing proper code, maintaining open-source software, and supporting users of our tools within and outside the lab. The practice of coding is a strong emphasis in the lab, but this is also an environment where trainees can grow and learn these skills, starting from various research backgrounds and levels of experience.
Lab members are encouraged to take the initiative in spinning out modular computational tools for their research when they may be relevant to the lab’s broader work. Shared tooling contributes substantially to lab infrastructure, and improves the robustness of the work we do. We support a shared ecosystem of tools in the lab and cultivate open and collaborative coding practice that prepare trainees for various career paths, within academia and industry.
Featured projects

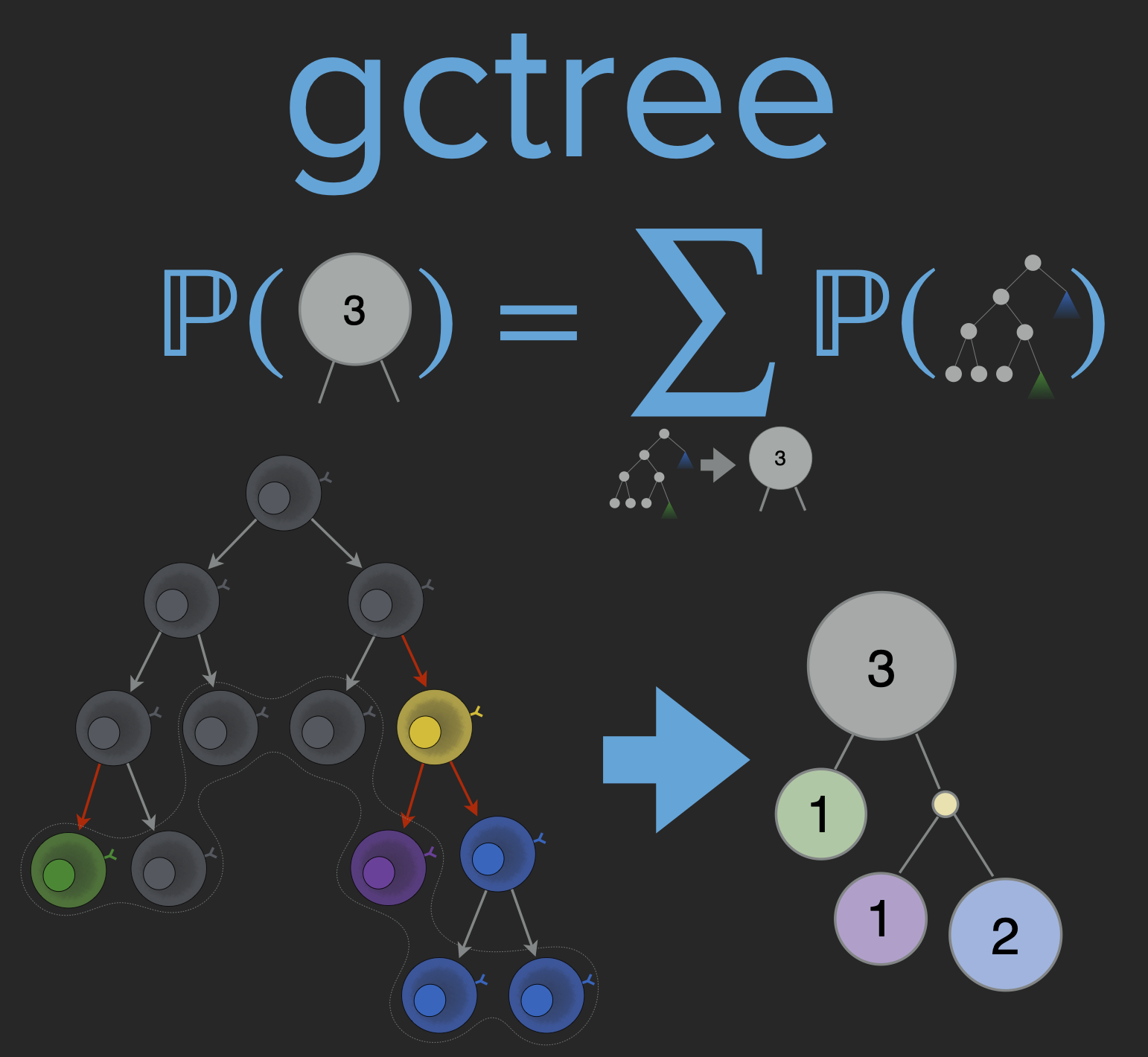
A Python package for tree simulation from general birth-death-mutation-sampling processes
multidms
A Python package that extends global epistasis models for joint estimation from deep mutational scans from multiple backgrounds

A Python package and command line utility for annotating the local ancestral sequence context of biallelic SNPs